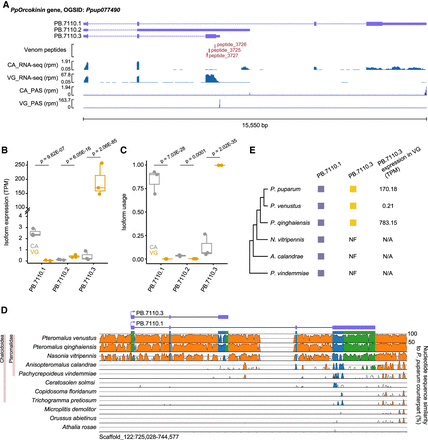
Evolutionary dynamics of the orcokinin gene and its venom isoform. (A) PpOrcokinin exhibits three isoforms (PB.7110.1, PB.7110.2, and PB.7110.3) expressed in the venom gland (VG) and carcass (CA). From top to bottom: isoforms, peptides supported by venom proteome, RNA-seq, and PAS-Seq. (rpm) reads per million. For supporting evidence of the venom PB.7110.3 protein, please refer to Supplemental Figure S26. (B) Expression levels of the three PpOrcokinin isoforms in VG and CA, displaying the adjusted P-values from the differential expression analysis conducted with DESeq2. (C) Isofrom usage of the three PpOrcokinin isoforms in VG and CA, showing the FDR (Benjamini–Hochberg) adjusted P-values from the differential isoform usage analysis. Isoform usage is calculated as expression of a specific isoform divided by the expression of the parent gene. (D) VISTA sequence conservation plot of the orcokinin gene in representative hymenopterans species, using P. puparum as the reference. Genomic sequences exhibiting over 50% sequence identity compared to the reference are depicted. Noncoding regions are indicated in orange, exons in blue, and UTRs in green. Conserved regions with at least 70% identity over a 100 bp window are shown in the corresponding color. The protein structures of two coding isoforms (PB.7110.1 and PB.7110.3) are displayed. A zoomed view of the exon3 alignment of PB.7110.3, including a specific DNA insertion in P. venustus, can be found in Supplemental Figure S27. (E) Schematic representation of orcokinin isoforms in six parasitoid wasps, presented to the right of the species tree. Filled boxes represent the presence of the corresponding isoform. (NF) not found. The isoform expression values (TPM) in VG are provided. (N/A) not applicable.











