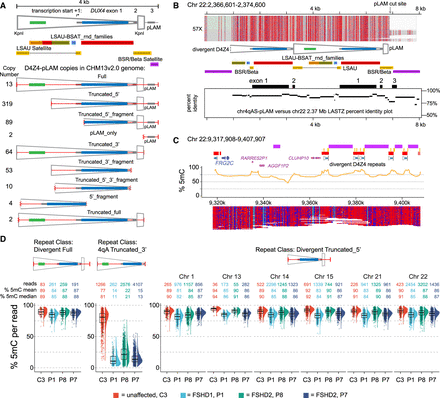
Methylation profiles of divergent D4Z4 repeats. (A) Copy number and structure of D4Z4-pLAM repeats found in the T2T CHM13 v2.0 genome (see Supplemental Table S4 for genomic locations and Fig. 1 for LSAU-BSAT composite annotations). (B) Methylation profile of a Chr 22 full-length, divergent D4Z4 repeat. The LASTZ percent identity (PIP) plot shows the distribution of sequence identity along the length of this repeat compared to the distal Chr 4qA-S D4Z4 repeat. (C) Methylation plots of reads mapping to Chr 22:9,317,908–9,717,807 region from the C4 control (R10.4.1_e8.2 nanopores). (D) Distributions of CpG methylation levels of divergent versus 4qA D4Z4 repeats. Each point represents the % 5mC of a single read within the region that aligns completely with the divergent repeat (Full or Truncated_5′) or the 3.3 kb 4qA D4Z4 repeat. The C3 sample was sequenced using R10.4.1_e8.2 nanopores.











