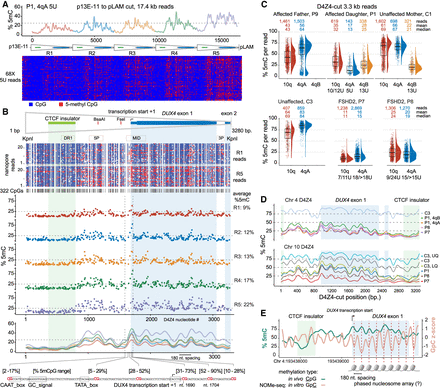
Fine-scale D4Z4 repeat methylation patterns. (A) Methylation frequency and modbamtools plots of the 4qA 5U reads from participant P1 using the “smooth_ksmooth” function (smoothness parameter = 10) and single-read plots using modbamtools. (B) An annotated methylartist plot of the 3.3 kb R1 and R5 repeats from a subset of reads in (A) with 322 CpG residues represented as blue (unmethylated) or red (methylated) dots. The middle panel shows the average methylation frequency for the CpGs aligned from R1 through R5 from the 5U reads in (A). The lower panel shows the smoothed methylation frequency plot for each repeat, with the expanded location of six CpG residues in the region containing the DUX4 proximal promoter and transcription start from Dixit et al. (AF117653.3). (C) Methylation levels of individual 3.3 kb D4Z4-cut reads with % 5mC per read on the y-axis, and annotated for 10q, 4qA, and 4qB read counts and their mean/median methylation levels. Upper plot = FSHD1 trio segregating a pathogenic 5U repeat; lower plot = unaffected versus FSHD2 participants. (D) Smoothed methylation frequency plots of D4Z4-cut reads from (C). (E) % 5mC and nucleosome occupancy and methylation sequencing (NOMe-seq) GpC Z-scores plotted for the second Chr 4 D4Z4 repeat (centromeric) using in vivo/in vitro methylation data from the HG002/NA24385 lymphoblastoid cell line.











