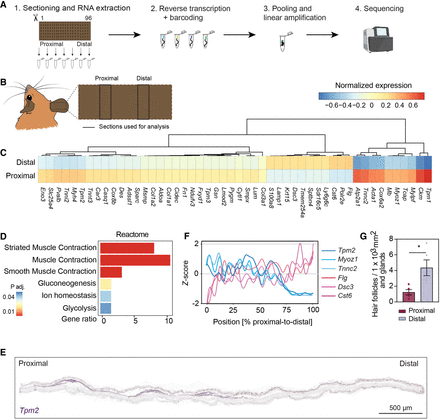
tomo-seq identifies asymmetric distribution of cell types across the Acomys ear. (A) Workflow, see text for details. Images were created with BioRender (https://www.biorender.com). (B) Schematic representation of the paired within-sample comparison on uninjured ears, proximal versus distal sections. (C) Paired within-sample comparison on uninjured ears, proximal versus distal. Shown are the top 50 differentially expressed genes (pad j <0.1, log2 fold change <−1 or >1). Color code indicates the averaged normalized expression per region (n = 2). (D) Enrichment analysis for significant genes at the proximal region using Reactome. Note that genes associated with the term “muscle” are strongly enriched in the proximal sections. (E) In situ hybridization on an ear section against Tpm2 mRNA, a muscle marker. Scale bar, 500 μm. (F) Line plot of Z-scored averaged gene expression patterns obtained by tomo-seq in the uninjured condition from the proximal to the distal side of the ear (n = 2). Shown are six representative genes selected from (C). (G) Barplot representing mean ± SEM of the counted hair follicles and glands in an uninjured ear proximal versus distal. (Paired t-test, [∗∗] P < 0.01; [∗] P < 0.05; n = 5.) (See also Supplemental Fig. S2; Supplemental Table S1.)











