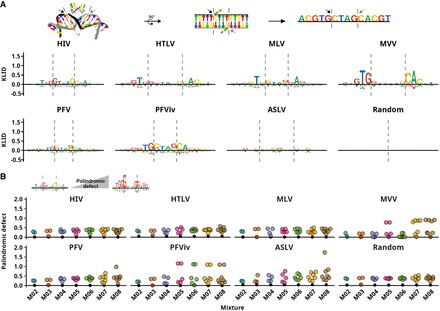
Identification of palindromic and asymmetric mixture components. (A) Scheme of retroviral integration and Kullback–Leibler information divergence (KLID)-based sequence logos representing complete sets of retroviral IS and a set of random genomic sequences. The arrows and the dashed lines mark the cleavage sites where the strand transfer reaction takes place. Logos represent IS sequences spanning 8 nt to each side from the center of the sequences. KLID values express the informativity of the nth position in sequence alignment relative to the global nucleotide frequencies. The character heights are proportional to the respective contributions of the nucleotides to the value of KLID. (B) Representation of palindromic defect that is defined as the dissimilarity between the PPM and its reverse-complement and is equal to zero for palindromic PPMs. All PPMs of retroviral IS component mixtures are represented. On the x-axis, mixtures are ordered by the number of components from two-component (M02) to eight-component (M08) mixture. The palindromic defect of the complete IS set (sequence logos in Fig. 1A) is included as a dark circle in each column.











