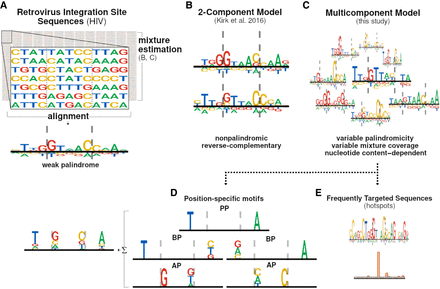
Origin of palindrome consensus motifs at sites of retroviral integration. Generally, a palindromic DNA motif composed of two reverse-complementary half-sites appears when palindromic sequences are aligned. (A) A weak palindromic motif appears when the sequences retrieved from the integration sites of retroviruses are aligned. The nature of the motif suggests the low frequency of the palindrome in aligned DNA sequences. Instead of whole-set alignments, mixture model estimates can be used to describe the possible submotifs in the sequence population. (B) A constrained two-component mixture model was used to analyze the consensus-forming submotifs, where an asymmetric motif appears on either of the half-sites of the target DNA (tDNA). (C) In this work, we used unconstrained multicomponent mixture models formed by at least two components (an eight-component mixture is depicted here). (D) Based on the submotifs appearing in multicomponent mixtures, we further performed quantitative analysis of the position-specific motifs. The major position-specific motifs observed include positional palindromes (PPs), broken palindromes (BPs), and asymmetric pairs (APs). (E) We described subpopulations of the frequently targeted sequences represented by low-abundant components in highly decomposed mixtures. We subsequently showed that one such component represents an abundant local hotspot of HIV-1 integration. Dashed lines in the sequence logos mark the cleavage sites of retroviral IN.











