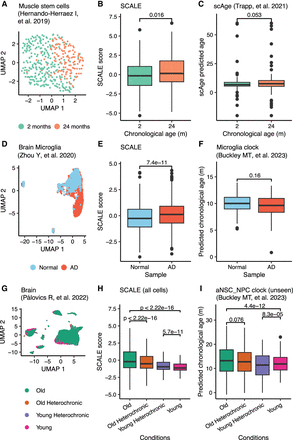
SCALE outperforms other single-cell aging clocks in assessing impacts of disease and rejuvenating interventions on aging. (A) UMAP plot of muscle stem cells. Different colors show different chronological ages. (B) SCALE scores of muscle stem cells (using single-cell transcriptomic profiles from single-cell M&T-seq). Data are presented as the mean ± SD. (C) ScAge predicted age of muscle stem cells (using single-cell DNA methylation data from single-cell M&T-seq). Data are presented as the mean ± SD. (D) UMAP plot of brain microglia cells from wild-type mice (normal; blue) and AD mouse models (AD; red). (E) SCALE scores of brain microglia cells from wild-type mice (normal; blue) and AD mouse models (AD; red). (F) Predicted chronological age of brain microglia cells by the method reported by Buckley et al. (2023). (G) UMAP plot of brain cells from old mice (green), young mice (magenta), old heterochronic mice (orange), and young heterochronic mice (purple). (H) SCALE scores of brain cells from parabiosis mice. P-values using a two-sided unpaired Student's t-test are shown. (I) Predicted chronological age of brain cell types (ependymal cell, macrophage, neuron, oligo pre cell, pericyte, and T cell) lacking clocks trained by Buckley et al. (2023). This box plot shows the results calculated by the aNSC_NPC clock, which incorrectly predicted cells from young heterochronic mice with lower ages than these of young mice and was not able to distinguish cells from old heterochronic mice and old mice, developed by Buckley et al. P-values using a two-sided unpaired Student's t-test are shown.











