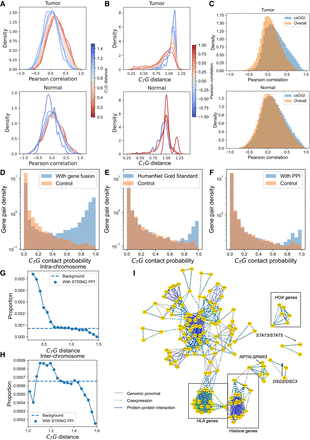
Passage of gene–gene interplay from genomic level to transcription and protein levels in colorectal cancer. (A) The distribution of transcriptional Pearson correlation under different CTG distance of the whole chromosome; the color of each line indicates the corresponding CTG distance. (B) The distribution of CTG distance under different Pearson correlation of the whole chromosome; the color of each line indicates corresponding Pearson correlation coefficient. (C) The distribution of correlation of gene pairs with csGGIs and overall background. (D) Distribution of CTG contact probability for gene pairs with (blue) and without (orange) gene fusion events reported in FusionGDB2 data set. (E) Distribution of CTG contact probability for gene pairs with (blue) and without (orange) similar biological process reported in HumanNet. (F) Distribution of CTG contact probability for gene pairs with (blue) and without (orange) PPIs reported in STRING database (G) The proportion of intra-chromosomal gene pairs with STRING PPI at different CTG distances in the tumor sample. The proportion refers to number of gene pairs with PPIs at fixed CTG distance/number of all gene pairs at fixed CTG distance. The background refers to number of gene pairs with PPIs/number of all gene pairs at all CTG distance. (H) The proportion of inter-chromosomal gene pairs with STRING PPI at different CTG distances in tumor sample. (I) The gene network integrates colon cancer–related gene–gene interplay at the DNA, RNA, and protein levels. The three kinds of edges indicate gene–gene interplays at three levels.











