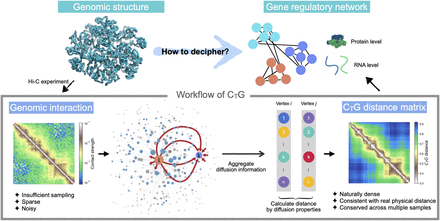
Schematic overview of CTG. The sparse Hi-C contact matrix is the starting data type of CTG. Taking the Hi-C contact matrix as the adjacency matrix of a graph, CTG uses a diffusion-based strategy to uncover the geometry of genomic structure from Hi-C data. CTG quantifies the diffusion property of each vertex by aggregating global diffusion information from the vertex to other vertices, respectively (as the red arrows illustrate). The CTG distance between pairwise vertices is calculated by the similarity of their diffusion properties. CTG allows for a genome-wide insight into deciphering the gene regulation information coded in genomic structure.











