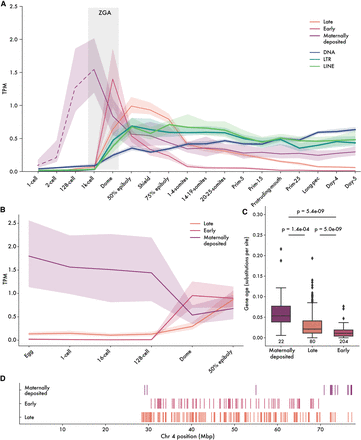
Embryonic expression of FZNF genes in zebrafish. (A) Median expression trajectory of 306 genes with expression of at least 0.5 TPM in at least one stage, separated by cluster. Shaded borders represent 90% confidence intervals. Light gray bar shows approximate timing of ZGA. Expression trajectory of maternally deposited FZNFs before the 1000-cell stage is dashed to reflect the fact that poly(A)-selected RNA sequencing is not an accurate reflection of true mRNA levels before ZGA. (B) FZNF expression from rRNA-depleted reads, before ZGA. (C) Maternally deposited genes are significantly older, whereas early-stage FZNFs are young and generally unique to D. rerio. Ages were quantified as branch lengths on a neighbor-joining phylogenetic tree generated from pairwise distances. (D) The majority of maternally deposited FiNZ genes are located in a specific, subtelomeric cluster. Each vertical line represents the midpoint of an expressed FZNF gene.











