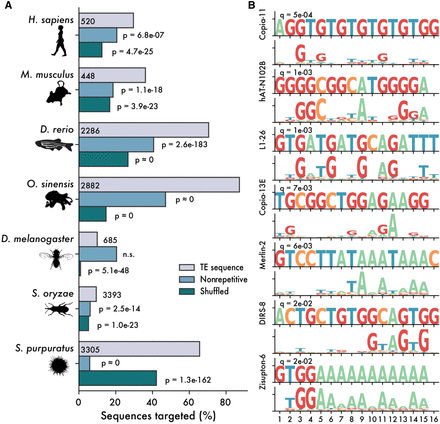
ZNFs are predicted to bind TE sequences. (A) Percentage of sequences matched by predicted ZNF motifs, with a Q-value cutoff of 0.05. Gray bars represent matches to libraries of consensus TE sequences, and blue bars represent matches in equally sized libraries of nonrepetitive genomic sequence. Green bars represent shuffled motifs, searched against consensus TE libraries. P-values are calculated as one-tailed binomial tests for the probability of observing at least x matches in the TE library, using binding frequency to nonrepetitive sequences or with shuffled motifs as a base probability. (B) Selected matches between Danio rerio consensus TE sequences and ZNF ORFs.











