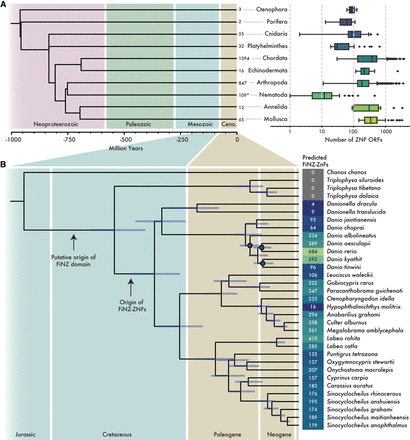
Annotation of ZNF ORFs and genes in metazoan genome assemblies. (A) Boxplots showing distributions of ZNF ORF counts per species in each major taxonomic phylum. Adjacent to boxplots are the number of representative species per phylum. (*) Number by nematodes excludes those with zero tandem repeat ZNF ORFs (defined as containing at least five repeats). Phylogeny was acquired from TimeTree5 (Kumar et al. 2022). (B) Maximum likelihood phylogenetic tree of Cypriniformes species with high-quality genome assemblies. Numbers represent counts of FZNF genes using improved gene annotations. Nodes marked in blue have lower than 95/95 support for Shimodaira–Hasegawa approximate likelihood ratio test/ultra-fast bootstraps, respectively.











