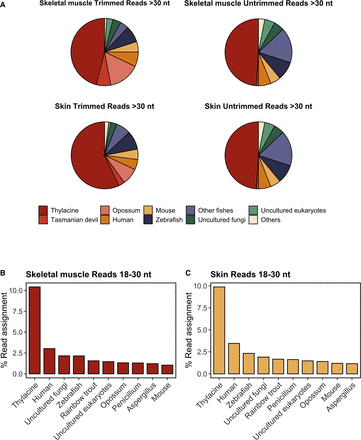
Metatranscriptomic analyses of thylacine skin and skeletal muscle samples. (A) Proportion of trimmed and untrimmed RNA reads (>30 nt) assigned using the KrakenUniq pipeline from thylacine skeletal muscle and skin tissues. Only species with more than 1000 k-mers and more than 200 species-specific reads are shown. Percentage of RNA reads aligned to the 10 most abundant species detected, focusing specifically on trimmed and untrimmed sequences uniquely mapped (MAPQ ≥ 1) of size ≥18 and ≤30 nt in skeletal muscle (B) and skin tissues (C). Reads with multiple-species ambiguous mapping were discarded for calculating the percentage of read assignment. Global alignment was performed using Bowtie 2 with flags ‐‐end-to-end and ‐‐very-sensitive.











