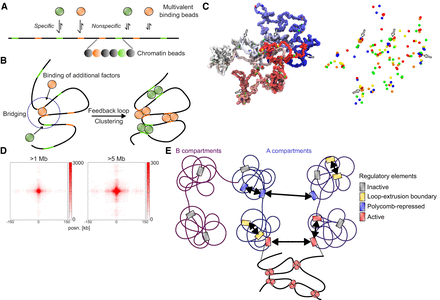
Simulations of chromatin and multivalent binding proteins. (A) Schematic depiction of the model with different multivalent binding beads (top) that bind specifically to colored binding sites and nonspecifically to other chromatin beads (bottom). (B) Schematic illustration of bridging-induced clustering. Binding of a multivalent factor leads to bridging and increased concentration of chromatin and binding sites in the vicinity of the bound site. This leads to increased likelihood for more factors to bind, causing a feedback loop and clustering. (C) Snapshot of a configuration in the model. Left side represents chromatin (polymer colored from red to blue), with specific binding sites represented by differently colored beads. Right side represents only the specific binding sites. Instances of clustering are highlighted with arrows. (D) Cumulative contacts in virtual Hi-C between binding sites in the simulations at distances >1 Mb (left) and >5 Mb (right). (E) Proposed model for interactions between CREs. Inactive genes in A or B compartments do not interact. In the A compartment, CREs overlapping loop extrusion boundaries interact via cohesin at short range, and polycomb-repressed CREs interact at short and long range. Active regulatory elements interact with each other at both short and long range, possibly through bridging-induced clustering











