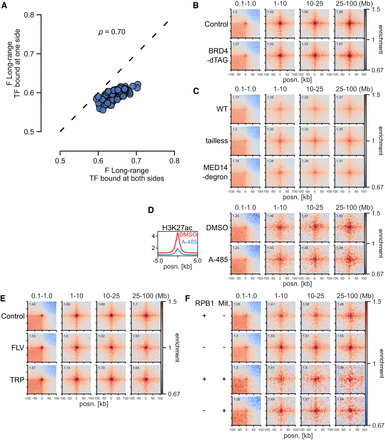
Independence of BRD4, Mediator, EP300 activity, and transcription. (A) Effect sizes of TFs for long-range (1- to 10-Mb) contact enrichment compared with TF-unbound CREs in mESCs. The x- and y-axes show enrichment for TF binding at both sides and only one side, but at least one other TF bound at the other side. (ρ) Spearman's correlation coefficient. (B) Pileup analysis between CGI Q4 regions in dTAG-BRD4 mESCs with or without BRD4. (C) Pileup analysis between CGI Q4 regions in WT, Med15−/− Med16−/− Med23−/− Med24−/− Med25−/− (tailless), and MED14− SMASh degron CH12 cells. (D, left) Mean H3K27ac signal in CGI Q4 regions in mESCs treated with DMSO or A-485. (Right) Pileup analysis between CGI Q4 regions in mESCs treated with DMSO or A-485. (E) Pileup analysis between CGI Q4 regions in Micro-C data from WT mESCs or treated with flavopiridol (FLV) or triptolide (TRP) for 45 min. (F) Pileup analysis between CGI Q4 regions in RPB1-AID mESCs with or without RPB1. The top two rows represent asynchronous cells; the bottom two rows represent cells arrested in mitosis (Mit +) and released into G1.











