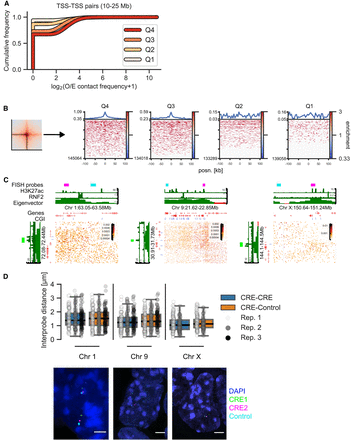
Distribution of contact frequencies and DNA FISH. (A) Empirical cumulative distribution frequency of O/E contact frequency values in the central 5-kb pixel between TSSs in different expression quartiles separated by 10–25 Mb. (B, left) Depiction of the “corner stripe” from a pileup. (Right) Heatmaps of O/E contact frequency values in individual corner stripes for TSS–TSS quartile interaction pairs at 10- to 25-Mb distance. The line plot on top represents the mean signal. (C) Hi-C matrices for regions on Chromosomes 1, 9, and X used in DNA FISH experiments. CGI marks CGIs without RNF2. (D, top) Interprobe distances measured by DNA FISH between distal CREs on Chromosomes 1, 9, and X or a CRE and an equidistant control region in the same A compartment. N = 289 (Chr 1), 292 (Chr 9), and 208 (Chr X) measurements each for CRE–Control and CRE–CRE. Boxplot lines denote maximum (excluding outliers), interquartile range (upper and lower bound of box), median (center of box), and minimum values. (Bottom) Example images of DNA FISH. Blue indicates DAPI; green, CRE1 probe; magenta, CRE2 probe; and cyan, control probe (adjacent to CRE2 in the genome). Scale bar, 2 μm.











