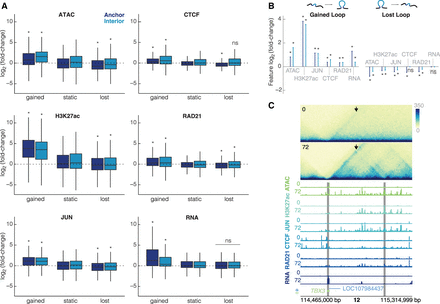
Changes in chromatin features are correlated with changes in looping. (A) Intersections of each feature at the loop anchors and interior for gained, static, and lost loops (dark blue = anchor, light blue = interior). All plots are on the same scale for the y-axis, showing log2(fold-change). Wilcoxon rank-sum test was performed for each feature to compare gained/lost anchors to static anchors and gained/lost interiors to static interiors; asterisks represent P < 0.05. (B) Median unshrunken log2(fold-change) of each data set at gained loops (left), and lost loops (right). Asterisks represent P < 0.05, dots represent P > 0.05. (C) 600-kb region around a loop at the TBX3 locus at 10-kb resolution. The arrow is pointing to the gained loop. Signal tracks for ATAC-seq, H3K27ac, JUN, CTCF, RAD21, and RNA for K562s at 0 h and differentiated megakaryocytes at 72 h show increased occupancy for all features. Gray bars indicate 10-kb loop anchors.











