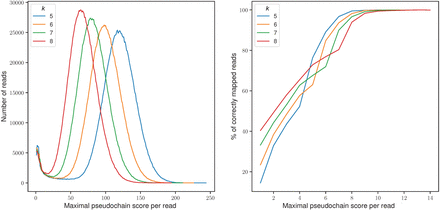
Effect of pseudochain score on mapping accuracy. The x-axis in both subfigures corresponds to the score of the maximal pseudochain per read; on the left, the y-axis denotes the total number of reads with the corresponding maximal pseudochain score; on the right, the percentage of reads that are correctly mapped (assessed by paftools mapeval) with the corresponding maximal pseudochain score. Only the scores below a threshold s, where s denotes the maximum score at which a read was mapped incorrectly, are plotted. The parameters used for mapquik were k = 5–8, ℓ = 31, δ = 0.01.











