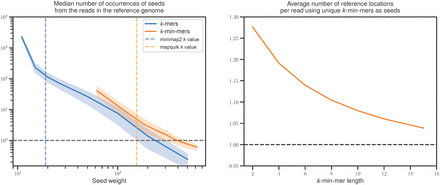
Increased sensitivity and specificity of k-min-mers versus long k-mers. Both panels use the human reference genome CHM13v2.0 and the HG002 DeepConsensus HiFi reads. (Left) Each continuous line indicates the median abundance of read k-mers (darker blue line) and k-min-mers (lighter orange line) in the reference, averaged across all reads (the closer to one, the better). The vertical dashed darker blue line (respectively, the lighter orange line) corresponds to the seed length chosen by minimap2 (respectively, by mapquik). The median is computed from a random subsample of 50,000 HG002 reads. (Right) Average number of reference genome locations indicated by seed matches for each read using k-min-mers (the closer to one, the better). k-min-mer parameters are ℓ = 31, δ = 0.01 with k = 2–10 (left) and 2–15 (right). Regular k-mer lengths are k = 12–500.











