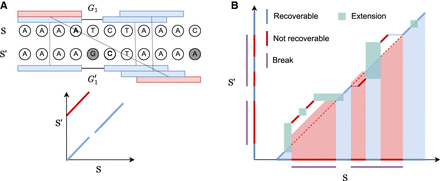
Seed-chain-extend visualized on an alignment matrix. Anchors are short diagonal matches in the matrix. Extension corresponds
to performing dynamic programming (DP) on the submatrix between anchors. (A) k-mer anchors under mutations and their corresponding alignment matrix. Blue anchors are homologous anchors, and red are spurious anchors. (B)  consists of anchors and DP matrices. Recoverable bases correspond to the bases on the homologous diagonal where a k-mer lies or is accessible by the green DP matrix between gaps. Breaks cover nonrecoverable sections.
consists of anchors and DP matrices. Recoverable bases correspond to the bases on the homologous diagonal where a k-mer lies or is accessible by the green DP matrix between gaps. Breaks cover nonrecoverable sections.











