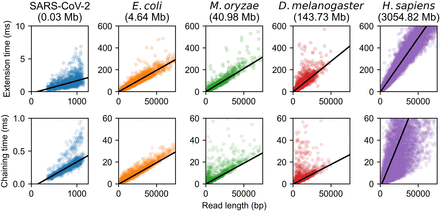
Extension and chaining time for real ONT reads with ≈5% sequence divergence. k-mer size was chosen to be k = C log n for reference length n and 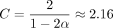 with error parameter θ = 0.05. Exact data sets are described in Supplemental Table S1. Note the scaled axes for the SARS-CoV-2 reads, which were much smaller than the other data sets. The well-fit linear regression lines indicate essentially
linear runtime in read length with fixed reference length n (i.e., fixed k = C log n and constant C), although larger human chaining time variance is owing to repetitive k-mers.
with error parameter θ = 0.05. Exact data sets are described in Supplemental Table S1. Note the scaled axes for the SARS-CoV-2 reads, which were much smaller than the other data sets. The well-fit linear regression lines indicate essentially
linear runtime in read length with fixed reference length n (i.e., fixed k = C log n and constant C), although larger human chaining time variance is owing to repetitive k-mers.











