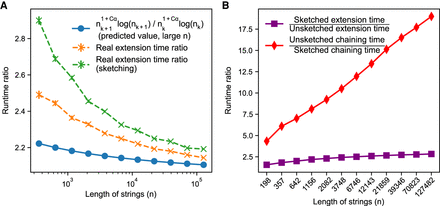
Runtime results on seed-chain-extend between for two sequences of length n and 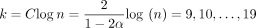 , where θ = 0.10 and α = −log 4(1 − θ). (A) The y-axis shows the runtime ratios (with 95% confidence intervals) for iteration k + 1 divided by runtime for iteration k, where the sequence length nk is plotted on the x-axis. As n grows, both the sketched and nonsketched extension ratios asymptotically approach the predicted ratio of
, where θ = 0.10 and α = −log 4(1 − θ). (A) The y-axis shows the runtime ratios (with 95% confidence intervals) for iteration k + 1 divided by runtime for iteration k, where the sequence length nk is plotted on the x-axis. As n grows, both the sketched and nonsketched extension ratios asymptotically approach the predicted ratio of 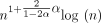 runtimes. (B) Multiplicative speed-up for sketched versus nonsketched alignment when sketching with density
runtimes. (B) Multiplicative speed-up for sketched versus nonsketched alignment when sketching with density 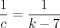 , where k = C log n. The slow-down for extension flattens out, whereas the chaining speed-up is almost linear on the log-scaled x-axis; chaining speed-up dominates extension slow-down as predicted by our theory.
, where k = C log n. The slow-down for extension flattens out, whereas the chaining speed-up is almost linear on the log-scaled x-axis; chaining speed-up dominates extension slow-down as predicted by our theory.











