Proving sequence aligners can guarantee accuracy in almost O(m log n) time through an average-case analysis of the seed-chain-extend heuristic
Abstract
Seed-chain-extend with k-mer seeds is a powerful heuristic technique for sequence alignment used by modern sequence aligners. Although effective in
practice for both runtime and accuracy, theoretical guarantees on the resulting alignment do not exist for seed-chain-extend.
In this work, we give the first rigorous bounds for the efficacy of seed-chain-extend with k-mers in expectation. Assume we are given a random nucleotide sequence of length ∼n that is indexed (or seeded) and a mutated substring of length ∼m ≤ n with mutation rate θ < 0.206. We prove that we can find a k = Θ(log n) for the k-mer size such that the expected runtime of seed-chain-extend under optimal linear-gap cost chaining and quadratic time gap
extension is O(mnf(θ) log n), where f(θ) < 2.43 · θ holds as a loose bound. The alignment also turns out to be good; we prove that more than 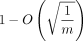 fraction of the homologous bases is recoverable under an optimal chain. We also show that our bounds work when k-mers are sketched, that is, only a subset of all k-mers is selected, and that sketching reduces chaining time without increasing alignment time or decreasing accuracy too much,
justifying the effectiveness of sketching as a practical speedup in sequence alignment. We verify our results in simulation
and on real noisy long-read data and show that our theoretical runtimes can predict real runtimes accurately. We conjecture
that our bounds can be improved further, and in particular, f(θ) can be further reduced.
fraction of the homologous bases is recoverable under an optimal chain. We also show that our bounds work when k-mers are sketched, that is, only a subset of all k-mers is selected, and that sketching reduces chaining time without increasing alignment time or decreasing accuracy too much,
justifying the effectiveness of sketching as a practical speedup in sequence alignment. We verify our results in simulation
and on real noisy long-read data and show that our theoretical runtimes can predict real runtimes accurately. We conjecture
that our bounds can be improved further, and in particular, f(θ) can be further reduced.
Footnotes
-
[Supplemental material is available for this article.]
-
Article published online before print. Article, supplemental material, and publication date are at https://www.genome.org/cgi/doi/10.1101/gr.277637.122.
-
Freely available online through the Genome Research Open Access option.
- Received January 3, 2023.
- Accepted March 16, 2023.
This article, published in Genome Research, is available under a Creative Commons License (Attribution-NonCommercial 4.0 International), as described at http://creativecommons.org/licenses/by-nc/4.0/.











