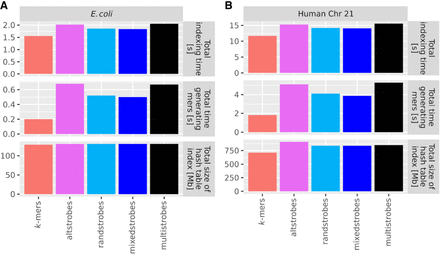
StrobeMap benchmarking. To benchmark our two new seeding techniques, Chromosome 21 of the GRCh38 human genome assembly (B) and one E. coli genome (assembly GCA 003018575.1 ASM301857v1) (A) were indexed with k-mers (k = 30), randstrobes (2, 15, 25, 50), mixedstrobes (2, 15, 25, 50, 0.8), altstrobes (2, 10, 20, 25, 50), and multistrobes (2, 5, 25, 25, 50) using StrobeMap. All experiments were repeated 10 times and the average (mean) was computed to account for variance in computer processing speed.











