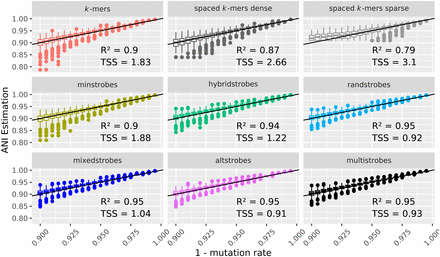
ANI-estimates of simulated sequences. ANI estimations on simulated data as estimated by Adjusted Mash distance (Methods) with k-mers, spaced k-mers, minstrobes, hybridstrobes, randstrobes (2,15,25,50), mixedstrobes (2,15,25,50,0.8), altstrobes (2,10,20,25,50), and multistrobes (2,5,25,25,50). For each method, the square of the Pearson correlation coefficient (R2, higher is better) and the total sum of squares (TSS; lower is better) between ANI estimation and true mutation rate is given.











