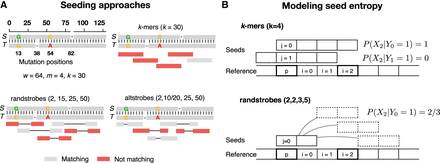
Sampling of k-mers, randstrobes, and altstrobes. Panel A shows two homologous sequences S and T containing four mutations with k-mers, randstrobes, and altstrobes sampled from T. When the sampled positions in a seed appear more random, it is likelier that at least one of the seeds in the region will sample only non-mutated positions with respect to S. Such non-mutated seeds are colored gray in the figure, and seeds containing mutated positions with respect to S are shown in red. Sampling non-mutated seeds in necessary to produce a seed match with seeds from S. Only a subset of five seeds in the region is shown for clarity. In the example, the seed constructs all sample 30 fixed positions in a window of w = 64 nucleotides which is the maximal span of a randstrobe with parameters (2,15,25,50). The mutations are distributed such that they destroy all shared k-mers in the region, and most of the randstrobes. Altstrobes have the possibility to sample k-mers of two different lengths at each site, which allow them a higher probability to match between mutations. Panel B illustrates our modeling of seed entropy for seed construct k-mers and randstrobes. In the case of k-mers, there is no pseudorandomness and therefore all probabilities are either 0 or 1, leading to an entropy of 0. Under uniform hashing, the randstrobes will have a probability of 2/3 of sampling position i owing to the three possible sampling positions of the second strobe. Boldfaced squares indicate the positions considered for the computation of probabilities shown in the figure.











