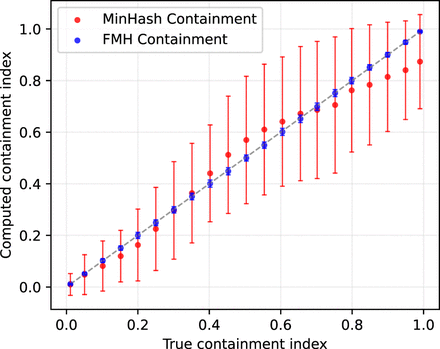
Figure 2.
True versus estimated containment index for both the Mash and FracMinHash approaches for two sets with dissimilar sizes. The containment index of a Staphylococcus genome is computed when C% of this genome is inserted into an assembled metagenome. Error bars indicate standard deviation over hash seed values.











