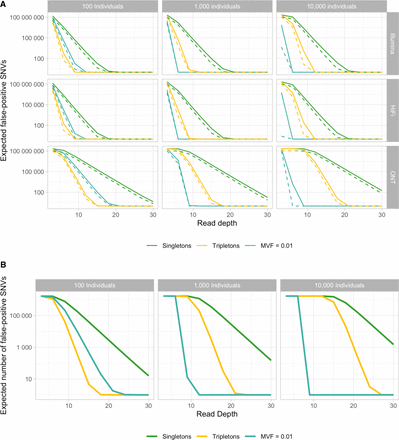
False-positive SNVs in non-B motifs. (A) Scenarios with different technologies corresponding to rows and numbers of diploid individuals (100, 1000, and 10,000) corresponding to columns. The average read depth per haploid genome is plotted on the x-axis, and the expected number of false-positive SNVs owing to errors is shown on the y-axis. Colors indicate different variant frequency filters (singletons, tripletons, and 1%) (for doubletons, see Supplemental Fig. S5), whereby the solid line corresponds to the cumulative number of all false-positive SNVs across non-B types; the dashed line, to the number of false-positive SNVs in an equally long stretch of B DNA. (B) Expected false-positive SNVs in middle guanines in guanine triplets at G4 motifs in Illumina sequencing. Within G4 motifs, there are 1,715,082 bp that fit the requirements of a middle guanine in a guanine triplet, which may have an extremely high error rate (Schirmer et al. 2016). Plotted are the numbers of expected false-positive SNVs, with the same coloring scheme and numbers of diploid individuals as in A.











