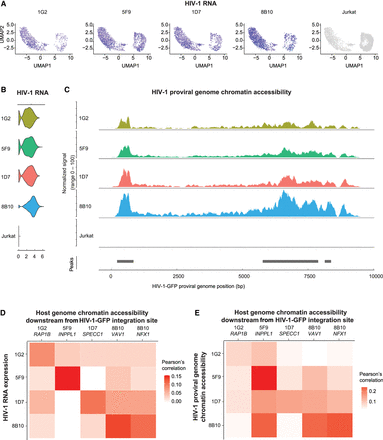
HIV-1 RNA expression correlates with HIV-1 proviral genome chromatin accessibility and host chromatin accessibility captured by single-cell DOGMA-seq. (A) UMAP dimensional reduction heatmap of HIV-1 RNA expression levels at the single-cell level in each HIV-1-infected cell line clone. (B) Log-normalized HIV-1 RNA expression level for each HIV-1-infected cell line clone and uninfected Jurkat clones. (C) Chromatin accessibility at the HIV-1 proviral genome across cell lines normalized to the total number of cells in each group by Signac. Peaks were identified by MACS2. (D) Pearson's correlations between HIV-1 RNA expression levels and host chromatin accessibility downstream from HIV-1 integration site across all cells within that clone. (E) Pearson's correlations between HIV-1 genome chromatin accessibility and host chromatin accessibility downstream from HIV-1 integration site across all cells within that clone. N = 11,333 cells total; 1314, 2869, 1853, 1906, and 3391 for 1G2, 5F9, 1D7, 8B10, and uninfected Jurkat cells, respectively.











