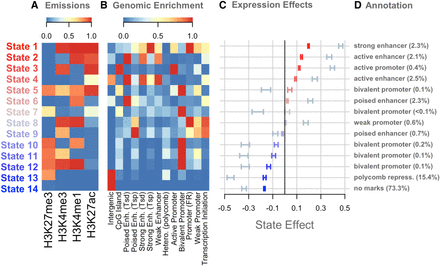
Overview of chromatin state composition, genomic distribution, and association with expression. (A) Emission probabilities for each histone modification in each chromatin state. Blue indicates the absence of the histone modification, and red indicates the presence of the modification. (B) The distribution of each state around functional elements in the genome. Red indicates that the state is enriched near the annotated functional element. Blue indicates that the state is depleted near the annotated functional element. Rows were scaled to run between zero and one for ease of visualization. Abbreviations are as follows: (Enh.) Enhancer, (Tsd) distal to the transcription start site, (Tsp) proximal to the transcription start site, (Hetero.) heterochromatin, (FR) flanking region. (C) The association between chromatin state variation and gene expression. Bars are colored based on the size and direction of the state's association with expression. Red/blue bars show the associations of chromatin state with gene expression across strains. Blue-gray bars show the associations of chromatin state with gene expression across genes. (D) Plausible annotations for each state based on genomic enrichments and association with gene expression. The numbers in parentheses indicate the percentage of the genome that was assigned to each state. (repress.) Repressor.











