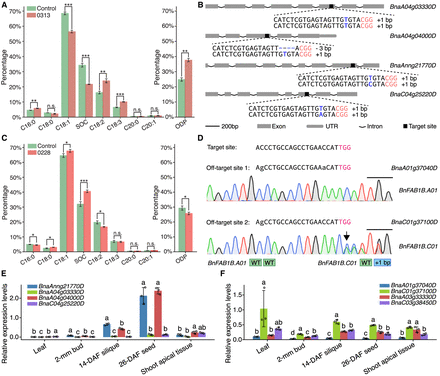
Significant alteration of the fatty acid profiles in two knockout mutants. (A) Fatty acid profile of the self-pollinated seeds in line 0313 of the T0 generation and the control background. (ODP) Oleic desaturation proportion, (SOC) seed oil content. (B) Detailed information for the editing interrogation of the four homoeologs of the BnEDA32 gene in the T0 generation. The PAM sequences are marked in red; the inserted nucleotides, in blue; and the deleted nucleotides, in blue dashes. The gene names are displayed on the right side of the structures. (C) Fatty acid profile of the self-pollinated seeds in the transgenic line 0228 of the T0 generation and the control background. (D) Editing interrogation in the two off-target sites of line 0228 in the T0 generation. The mismatched nucleotides are marked in lowercase, and the PAM sequences are marked in red and indicated by the black lines above the sequencing chromatograms. The arrow indicates the insertion site of the BnFAB1B.C01 gene. (E) Relative expression levels of the four homoeologs of the BnEDA32 gene in the five tissues of J9709 WT. (F) Relative expression levels of the four homoeologs of the BnFAB1B gene in the five tissues of J9709 WT. In A and C, the Student's t-test was used for statistical analysis: (n.s.) not significant, (*) P < 0.01, (**) P < 0.001, and (***) P < 0.0001. In E and F, different letters represent significant differences at P < 0.05 by two-way ANOVA with Tukey's HSD. (A,C,E,F) Error bars represent SDs.











