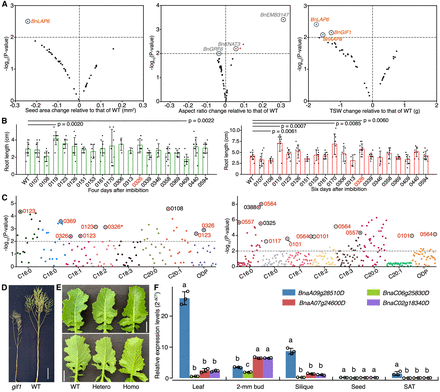
Phenotyping of the selected positive transgenic lines after interrogating the editing. (A) Phenotyping of seed-related traits. Each dot represents a transgenic line, and the corresponding edited gene is alongside. (TSW) Thousand-seed weight. (B) Phenotyping of root length at the stages of 4 and 6 d after seed imbibition. P-values less than 0.01 are shown. The ID of the plant with four edited homoeologs of the BnKNAT3 gene is marked in red. (C) Phenotyping of the fatty acid composition in self-pollinated (by bagging; left panel) and open-pollinated (naturally mature; right panel) seeds. Each dot represents a transgenic plant, and the mutant IDs are marked. (ODP) Oleic desaturation proportion. The asterisk indicates the ID of the plant with four edited homoeologs of the BnKNAT3 gene. Seeds used for the phenotyping in A–C were from plants of the T0 generation. (D) A representative photo displays the dwarf phenotype of the gif1 mutant in the T1 generation. Scale bar, 15 cm. (E) Representative photo shows the glossy phenotype of the examined kcs6.A09 homozygous mutant in the T1 generation. (Hetero) Heterozygote, (Homo) homozygote. Scale bar, 2 cm. (F) Relative expression levels of the four homoeologs of the BnKCS6 gene in the indicated five tissues of J9709 WT. (SAT) Shoot apical tissue. Different letters represent significant differences at P < 0.05 by two-way ANOVA with Tukey's honest significant difference (HSD). In A–C, the Student's t-test was used for the statistical analysis.











