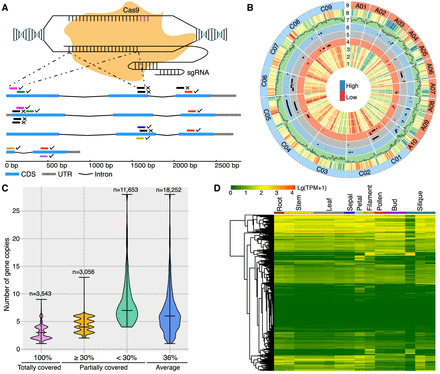
sgRNA design and characteristics of the selected 20-bp target sites. (A) The schematic diagram illustrates the three principles of sgRNA design. Each small bar near the corresponding target gene represents a 20-bp target site, and the identical ones are marked in the same full color. Ticks alongside the target sites indicate the selection, whereas crosses are on the opposite. This example displays the four homoeologs of the BnGPAT4 gene. (B) Circos plot visualizes the integrated information for the 18,414 selected target sites and the corresponding 10,480 target genes. (Tracks 1–3) Expression levels of the target genes in seeds 42 d after flowering (DAF; 1), 30-DAF seeds (2), and 17-DAF seeds (3), detected in our transcriptome data. (Tracks 4–6) Distribution of quantitative trait loci (QTLs) related to thousand-seed weight (TSW; 4), silique length (5), and seed oil content (6). (Tracks 7,8) Density distribution of the target sites (7) and the target genes (8). (Track 9) Nineteen chromosomes of the reference genome. (C) Density distribution of the target sites across the number of homoeologs. The widest part of the violin plot stands for the highest density of target sites; the center line is the median; and the top and bottom lines represent the entire data range (one group containing 65 homoeologs was removed from this analysis). An example for interpretation of this plot is as follows: Each of the 3543 target sites covers all homoeologs from the same homoeologous group and so on for the rest. (D) Expression features of the representative target genes (n = 7791) in different tissues of the ZS11 cultivar. Among the 7791 genes, 6220 (79.8%) were identified as expressed (with TPM ≥ 1) in at least one of the 16 tissues; however, expression of the last 1571 genes (20.2%) was not detected in any tissue.











