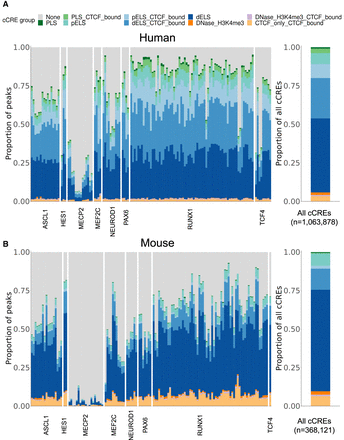
Figure 3.
Overlap of ChIP-seq peaks with annotated regulatory elements. (A, left) The proportion of peaks for each human ChIP-seq experiment, grouped by TR, that overlapped with a candidate cis-regulatory element (cCRE). (PLS) Promoter-like sequence, (ELS) enhancer-like sequence, (p) TSS proximal, (d) TSS distal. (Right) The proportional breakdown of all cCRE groups. (B) Same as in A, except for mouse ChIP-seq experiments.











