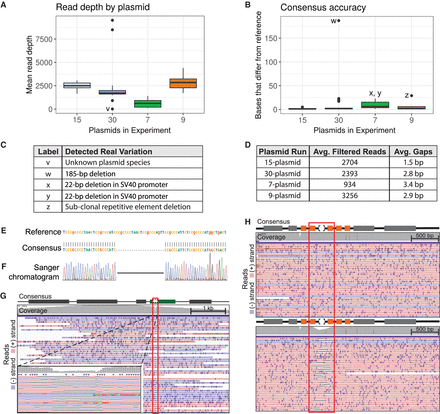
Real plasmid sequencing experiment characteristics and variant detection. (A) Per-plasmid read depth across pooled sequencing runs. (B) Per-plasmid count of bases in consensus sequence that differ from reference (gaps). (C) Table describing variation contributing to outliers (labeled points) in A and B. (D) Table summarizing read and gap data for experiments shown in A and B (gap counts do not include variants listed in C). (E) OnRamp alignment results showing a 22-bp deletion. (F) Sanger validation of deletion in E. (G) IGV browser view of reads mapping to deletion (red outline) from E in an SV40 promoter (green box). (Left inset) Zoomed view. (H) IGV view of reads mapping to a clone without (top) or a clone with (bottom) a subclonal repetitive element (orange boxes) deletion (red outline). IGV: Black lines are deletions; dark purple marks are insertions.











