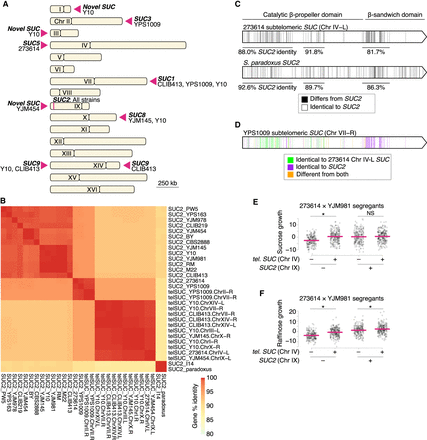
Subtelomeric copies of genes encoding invertase. (A) Invertase genes in the 16 genomes, with subtelomeric SUC loci and the SUC2 locus marked with wedges and a vertical bar, respectively. The SUC9 locus on Chr XIV (Naumov and Naumova 2010b) was not mapped to a specific arm, so we labeled both arms as SUC9. We did not observe SUC genes on Chr VIII, XIII, or XVI (SUC7, SUC4, and SUC10, respectively) (Naumova et al. 2014). (B) Gene percentage identity between SUC genes. (C) Gene sequence comparison of SUC2 from strain BY to the representative subtelomeric SUC gene from strain 273614 (top) and SUC2 from S. paradoxus (bottom). (D) Gene sequence comparison of the Chr VII SUC gene from strain YPS1009 to the subtelomeric SUC gene from strain 273614 and SUC2 from strain BY. (E,F) Scatterplots of growth on 2% sucrose or raffinose, respectively, of segregants of the 273614 × YJM981 cross, partitioned by SUC genotype at Chr IV and IX, as in Figure 3. Mean phenotypes are shown with magenta lines. (*) P < 0.05, one-tailed Student's t-test, with the alternative hypothesis that the extra SUC copy improves growth.











