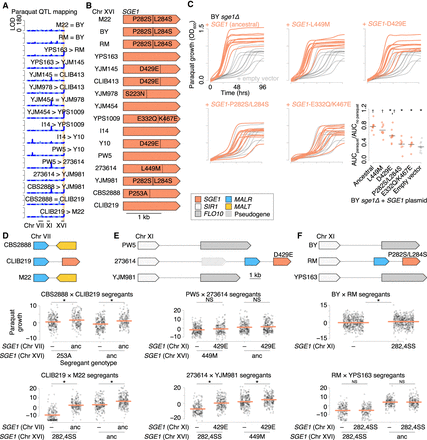
Diversity in SGE1 underlies widespread variation in paraquat tolerance. (A) QTL mapping of growth on 0.75 mM paraquat (Bloom et al. 2019), as in Figure 2A. The site of a recurrent QTL on Chr XVI is highlighted in peach. (B) Diagram of the SGE1 alleles on Chr XVI, with derived nonsynonymous variants labeled. (C) Growth over time in liquid YPD media with 3.5 mM paraquat for BY sge1Δ with plasmids carrying SGE1 from strains BY, YJM145, YPS1009, I14, or 273614 (orange) or empty vector (gray). n = 10 biological replicates. The last panel quantifies strain growth as the AUC for growth in paraquat normalized to the AUC for growth without paraquat (Supplemental Fig. S13). SGE1 alleles from strains BY, YJM145, and YPS1009 provided significantly less protection than the ancestral allele from strain I14. (*) P < 0.005, Tukey's HSD following one-way ANOVA. In addition, SGE1 alleles from strains YJM145, I14, and 273614 provided significantly greater protection than empty vector ([†] P < 0.001, Tukey's HSD), whereas alleles from strains BY and YPS1009 did not. (D–F) Additional SGE1 genes were found in strains CLIB219, 273614, and RM, respectively. Scatterplots below show growth on paraquat for segregants in crosses involving those strains, partitioned by SGE1 genotype. Plots are shown as in Figure 3, with normalization to growth on plates without paraquat. Mean phenotypes are shown with orange lines. (*) P < 0.05, one-tailed Student's t-test, with the alternative hypothesis that the extra SGE1 copy improves growth on paraquat.











