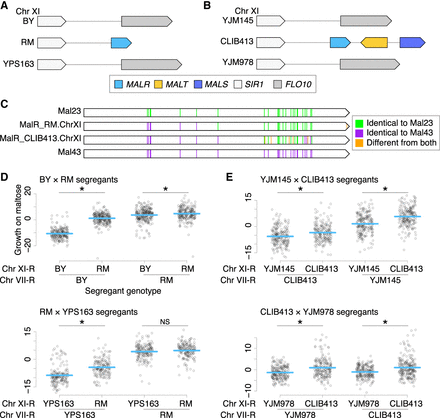
Maltose QTLs on Chr XI are caused by nonreference MAL loci. (A,B) Diagrams of MAL loci on Chr XI of RM and CLIB413, respectively, along with the MAL-less loci in the strains they were crossed to for QTL mapping. (C) Amino acid sequence comparison of Chr XI MALRs from RM and CLIB413 to Mal23 and Mal43. (D,E) Scatterplots of maltose growth of segregants from crosses involving strains RM and CLIB413, respectively, partitioned by segregant genotype at Chr XI and VII. Colony radiuses were measured by Bloom et al. (2019) after 48-h growth on maltose plates, normalized to colony radiuses on glucose plates, and then scaled to have a mean of zero and a variance of one. The mean phenotype of each genotypic grouping of segregants is shown with a blue line. (NS) Not significant, (*) P < 0.05, one-tailed Student's t-test, with the alternative hypothesis that the extra MAL genes improve maltose growth.











