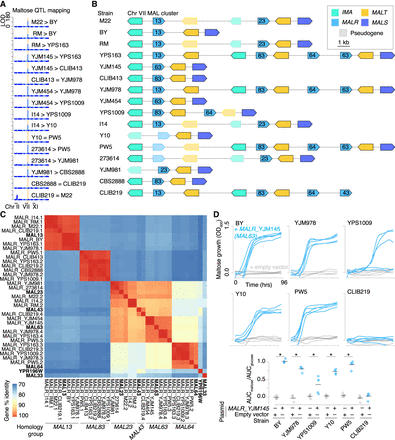
Diverse, highly complex MAL loci on Chr VII. (A) QTL mapping of maltose growth in 16 biparental S. cerevisiae crosses (Bloom et al. 2019). Each plot shows linkage (calculated logarithm of the odds [LOD] score), in one biparental cross, between segregant genotype and growth on 2% maltose, plotted against genomic coordinates. Twelve of 16 crosses have a QTL with LOD score greater than 35 at the right end of Chr VII, highlighted in peach; plots are labeled with which parent provided the Chr VII allele associated with superior maltose growth, if any. (B) Diagram of the Chr VII MAL locus in each strain. MALR genes are labeled by homology group, as determined in panel C. (C) DNA percentage identity between MALR genes found on Chr VII (McWilliam et al. 2013). Previously identified MALR genes are included for reference, in bold (Gibson et al. 1997). MALRs with >97% sequence identity are sorted into homology groups, labeled at the bottom. As seen by Song et al. (2015), we observed a homology group of MALRs with low sequence identity to any named MALR, which we labeled MAL83 (assigned systematic name YSC0072 at the Saccharomyces Genome Database [SGD; https://www.yeastgenome.org/]). (D, top) Maltose growth for maltose-poor strains BY, YJM978, YPS1009, Y10, PW5, and CLIB219 with plasmids carrying MALRYJM145 (blue) or empty vector (gray). n = 5 biological replicates, except for YJM978 + empty vector, which had four biological replicates. (Bottom) Growth was quantified as the area under the growth curve (AUC) for maltose normalized to the AUC for glucose (Supplemental Fig. S10). Maltose growth was significantly higher with MALRYJM145 than empty vector for strains BY, YJM978, YPS1009, Y10, and PW5. (*) P < 0.005, Tukey's HSD following one-way ANOVA.











