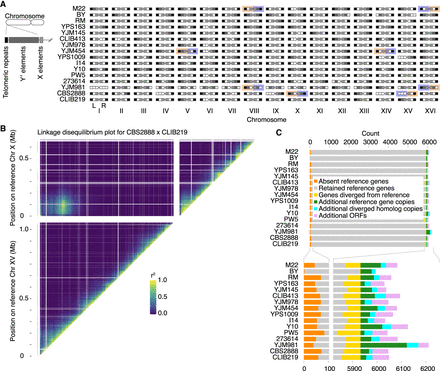
Long-read genomes. (A) Detection of telomeric TG1–3 repeats and telomere-associated Y′ or X element sequences at ends of assembled contigs. TG1–3 repeats were detected as more than 20 TG dinucleotides in the final 100 bp of a contig, whereas Y′ and X elements were detected by Liftoff. Dark, medium, and light gray shadings indicate the presence of TG1–3 repeats, Y′ elements, or X elements on a contig end, respectively, whereas white indicates the feature was not detected. Orange and purple highlighting mark chromosome arms affected by reciprocal translocations. (B) Linkage analysis of markers on reference Chr X and XV in the CBS2888 × CLIB219 cross. The existence of strong cross-chromosome correlation between Chr X and XV markers is consistent with a translocation between these chromosomes in strain CBS2888. (C) Gene and ORF content in assembled genomes. Genes and gene copies with >30% sequence identity to reference genes were classified as retained reference genes or diverged homologs using a threshold of 95% sequence identity to the reference gene.











