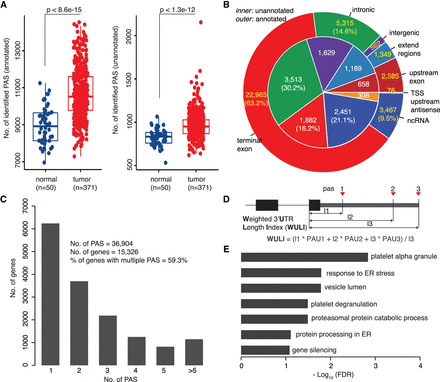
Applying APAIQ on a large-scale RNA-seq data set identifies tumor-associated APA events. (A) Boxplot showing the number of annotated and novel PASs identified by APAIQ in each RNA-seq sample from TCGA-LIHC. P-values were derived from Wilcoxon test. (B) Proportion of the identified PASs from a different genomic category. The outer circle indicates the identified PASs that are overlapped with the annotation and the inner pie presents the identified PASs that are not overlapped with the annotation. (C) Numbers of genes with different numbers of the identified PASs. (D) A schematic illustration of the calculation of weighted 3′ UTR length index (WULI). PAU stands for PAS usage and the l1, l2, and l3 represent the length from start to the terminal exon to each PAS. (E) Barplot showing the BH-adjusted P-value (FDR) of Gene Ontology terms and KEGG pathways enriched for genes with 3′ UTR shortening in tumor compared to normal.











