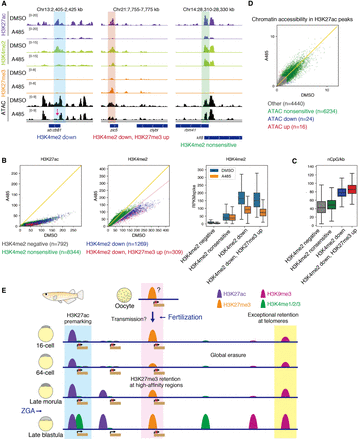
H3K27ac is required for proper histone modification patterning in late blastula embryos. (A) Track views showing the histone modification enrichments after spike-in normalization and chromatin accessibility in DMSO- or A485-treated embryos. The magenta arrow indicates where chromatin accessibility was altered after A485 treatment. (B) Comparison of H3K27ac enrichment and H3K4me2 enrichment in H3K27ac peaks between A485 and DMSO treatment by scatter plots (left) and boxplots (right). Yellow and pink lines indicate Y = X and Y = 0.8 × X−20, respectively. (C) Comparison of CpG density (number of CpGs/kb) in H3K27ac peaks between A485 and DMSO treatment by a boxplot (right). (D) Scatter plot showing chromatin accessibility in H3K27ac peaks in DMSO and A485 treatment. “Other” indicates the H3K27ac peaks with low ATAC-seq signals. Yellow line indicates Y = X. (E) Summary of histone modification reprogramming in medaka. All histone modifications more or less undergo erasure after fertilization, but some modifications escape complete erasure in medaka. H3K27ac premarks promoters and is required for proper patterning of histone modification (e.g., H3K4me2). H3K27me3 is probably transmitted maternally and retained at Polycomb high-affinity regions, although the enrichment was very limited. H3K9me3 at telomeric regions escapes complete erasure and is required for maintenance of chromosome stability during cleavage stage. On the other hand, H3K4 methylations and H3K9me3 except at telomeric regions are extensively erased after fertilization.











