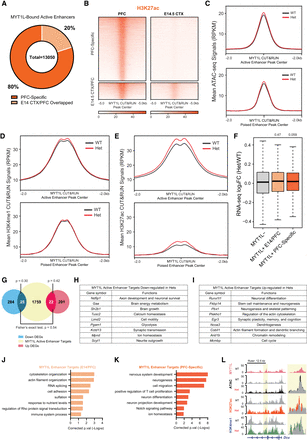
MYT1L suppresses enhancers that regulate neuronal migration and neuron projection development. (A) Majority of MYT1L-bound active enhancers were PFC-specific. (B) Heatmaps of MYT1L-bound active enhancers in PFC. (C) MYT1L loss increased its bound active enhancers but not poised enhancers’ chromatin accessibility. (D) MYT1L loss increased H3K4me1 levels at MYT1L-bound active enhancers but did not significantly increase H3K4me1 levels at MYT1L-bound poised enhancers. (E) MYT1L loss increased both its bound active and poised enhancers’ H3K27ac levels. (F) MYT1L active enhancer target genes showed increased expression in Het PFC. (G) Venn diagram showing the overlaps among dDEGs, MYT1L active enhancer targets, and uDEGs. There were more overlaps between MYT1L active enhancer targets and uDEGs than dDEGs (P = 0.01). (H) Functions of MYT1L active enhancer targets showing down-regulation in RNA-seq. (I) Functions of MYT1L active enhancer targets showing up-regulation in RNA-seq. (J) GO analysis on genes associated with MYT1L-bound (MYT1L+) E14 CTX/PFC overlapped active enhancers. (K) GO analysis on genes associated with MYT1L+ PFC-specific active enhancers. (L) Representative genome browser track showed MYT1L-bound Dcx active enhancer has higher ATAC-seq signals, H3K4me1 and H3K27ac levels in Het PFC than WT.











