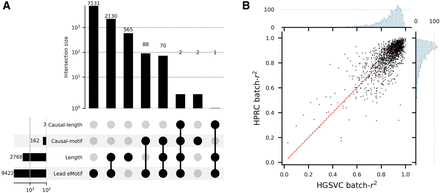
Fine-mapping of VNTR variants. (A) VNTR-centric view of gene-level eQTL discoveries and fine-mapping. Lead eMotif denotes any VNTRs that are associated with gene expression through at least one eMotif under gene-level discoveries. Length denotes any VNTRs that are associated with gene expression through length under gene-level discoveries. Causal-motif denotes any VNTRs for which a motif passes the fine-mapping procedure with a posterior inclusion probability ≥ 0.8 while being a significant eQTL under genome-wide P-value cutoff. Motifs with the lowest P-value for each VNTR–gene pair are included in the fine-mapping model in addition to VNTR length. (B) The batch-r2 of likely causal motifs on the HGSVC and HPRC data sets. Each bin on the marginal histogram spans 0.01.











