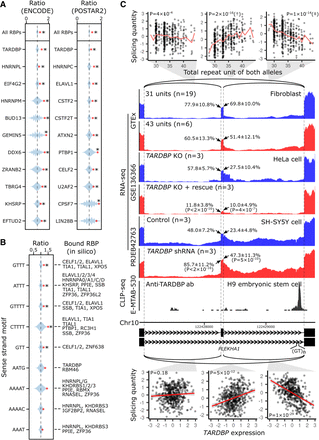
Mechanisms underlying spl-TRs. (A,B) Shown from top to bottom are the most enriched RBPs (A) or repeat motifs (B) among intronic spl-TRs (n = 5660). The count of each RBP or repeat motif observed in the 5660 intronic sTRs is shown by the red point against blue violin plots representing the kernel density estimation of counts in sets (n = 10,000 and 100,000 for repeat motifs and RBPs, respectively) of 5660 randomly selected intronic TRs among all the intronic TRs (n = 18,139). The counts are represented as the ratio to the median of the negative control sets. For each repeat motif, RBPs predicted to bind it by SpliceAid or ATtRACT are shown on the right (B). (*) Q-value < 0.1. (C) The spl-TR at PLEKHA1. Shown are correlations between TR dosages and splicing quantities in GTEx data (top panel), RNA-seq depth of samples with a smaller or larger TR dosage in GTEx data (second panel from the top), TARDBP-knockout HeLa cell lines rescued by exogenously re-expressing TARDBP or not (third panel from the top), and SH-SY5Y cells stably expressing a shRNA for TARDBP or control SH-SY5Y cells (fourth panel from the top), CLIP-seq depth of H9 embryonic stem cells using an anti-TARDBP antibody (fifth panel from the top), exon–intron structures (sixth panel from the top), and correlations between TARDBP expression and splicing quantities in GTEx data (bottom). These panels are drawn as in Figure 2C and D. (Third and fourth panels from the top) The splice site usages were compared between the two cell types using binomial generalized linear model, and the resulting P-values are shown in brackets. (Bottom) The significance of linear regression is given in the plots (threshold: 3 × 10−4). The red line and gray shading represent the regression line and 95% confidence interval, respectively. The expression and splicing levels are normalized (see Methods subsection “Correlation between RBP expression level and splicing quantity”).











