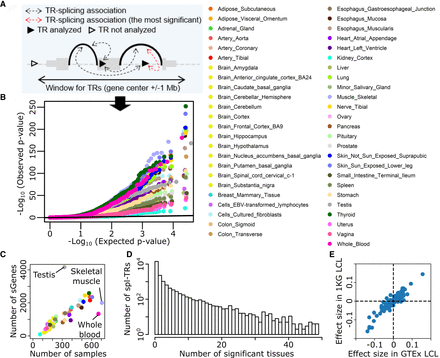
Discovery of spl-TRs across 49 tissues. (A) Selection of the most significant TR–splicing association in each gene. (B) QQ plot of P-values obtained from spl-TR mapping. Dots represent the most significant TR–splicing association in each gene. The black y = x line corresponds to the null hypothesis. (C) Number of associations per tissue as a function of tissue sample size. Tissue color codes are shown in B. (D) Sharing of TR–splicing associations across tissues. The x-axis: the number of tissues sharing a given association; the y-axis: the number of associations shared across a given number of tissues. (E) Comparison of effect sizes of 563 TR–splicing pairs between GTEx and 1 KG LCL data sets. Dots denote TR–splicing pairs.











