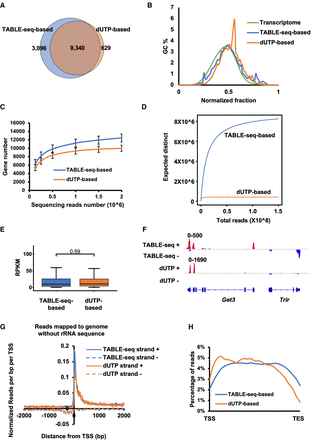
Benchmark between TABLE-seq-based and dUTP-based strand-specific RNA sequencing. (A) Venn diagram showing the overall detected genes between libraries generated by TABLE-seq-based and dUTP-based methods. (B) Percentage of GC content in TABLE-seq-based and dUTP-based methods. (C) TABLE-seq-based and dUTP-based methods shared a plateau of saturation at around 1 million total reads. Saturation curves were constructed with random gradients. N = 2 M, 1.5 M, 1 M, 0.5 M, and 0.25 M reads from total reads. Data were mean ± SD. (D) The library complexity of TABLE-seq-based and dUTP-based methods. (E) Boxplot showing the read counts per gene in libraries generated by the indicated methods. (F) IGV views showing the enrichments of mapped reads at the sense (strand +) and antisense (strand −) transcripts. (G) Comparison of sequencing signals between libraries generated by the indicated methods. Reads were mapped to the genome without ribosomal RNA sequences. Sense (strand +) and antisense (strand −) transcripts associated with TSS are shown. (H) The reads distributions spanning the gene regions. (TES) Transcription end site.











