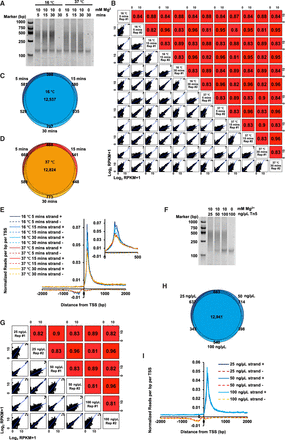
Analyses of tagmentation conditions in strand-specific RNA sequencing. (A) PCR results of TABLE-seq-constructed libraries at the indicated time and temperature are shown. cDNAs were tagmented at the indicated time and temperature, PCR amplified, and analyzed by TBE gel. Mg2+ was required for the enzymatic activity of Tn5 transposon. Samples treated without Mg2+, which inhibited the Tn5 transposon activity, were used as negative controls. (B) The Pearson's correlations among libraries generated by the indicated reaction time and temperature. The scatter plots show all detected genes. (RPKM) Reads per kilobase of exon per million reads mapped. (C) Venn diagram showing the overall detected genes among libraries generated by reactions at 16°C with the indicated reaction time. (D) Same as C, except libraries generated by reactions at 37°C are shown. (E) Comparison of sequencing signals among libraries generated by the indicated time and temperature. Sense (strand +) and antisense (strand −) transcripts associated with TSS are shown. The distributions at the first 500 bp are enlarged. (F) PCR results of TABLE-seq-constructed libraries by different amounts of Tn5 are shown. cDNAs were tagmented with the indicated amounts of Tn5, PCR amplified, and analyzed by TBE gel. Mg2+ was required for the enzymatic activity of Tn5 transposon. Samples treated without Mg2+, which inhibited the Tn5 transposon activity, were used as negative controls. (G) The Pearson's correlations among libraries generated by the indicated amounts of Tn5. The scatter plots show all detected genes. (H) Venn diagram showing the overall detected genes among libraries generated by the indicated amounts of Tn5. (I) Comparison of sequencing signals between libraries generated by the indicated amounts of Tn5. Sense (strand +) and antisense (strand −) transcripts associated with TSS are shown.











