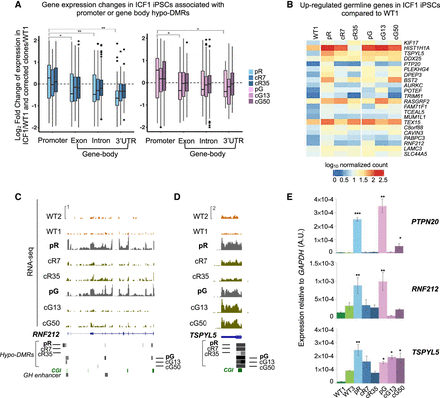
Transcriptional deregulation associated with promoter and intragenic hypo-DMRs. (A) Boxplot representation of the log2 fold change of the differentially expressed genes (pp > 0.8) with hypo-DMRs associated with regulatory elements (CGIs and GH promoters/enhancers) in pR (n = 471) and pG (n = 332) iPSCs and their corrected clones compared to WT1 iPSCs. The x-axis shows the gene feature category, and the y-axis denotes the log2FC of the genes in each ICF1 iPSCs versus WT1 iPSCs. The statistical significance of the differences in the log2FC of genes with hypo-DMRs annotated to promoters compared with genes containing hypo-DMRs annotated to other gene features was calculated using a nonparametric two samples two-sided Wilcoxon test with BH-FDR correction: (*) FDR < 0.01; (**) FDR < 0.001. (B) Heatmap representing the expression levels of germline-specific genes hypomethylated and up-regulated in ICF1 iPSCs. The scale denotes the log10 of normalized counts of genes compared between ICF1 iPSCs and corrected iPSCs to WT1 iPSCs. (C,D) Genome browser views of the expression profiles of the hypomethylated germline-specific genes RNF212 and TSPYL5. The coverage tracks display the RNA-seq expression level of the genes in WT1 and WT2 internal and external controls, ICF1, and corrected iPSCs. Gray boxes represent hypo-DMRs in ICF1 and corrected iPSC clones compared to WT1 and WT2. (E) RT-qPCR of the up-regulated RNF212, PTPN20, TSPYL5 germline genes in WT, ICF1, and corrected iPSCs. Each bar represents the mean of the relative expression compared to the expression of GAPDH in the same sample. The RT-qPCR data are presented as mean ± SD from independent triplicates, each amplified twice. Statistical analyses were performed using a one-tail two-sample Student's t-test compared to WT: (*) P-value < 0.05, (**) P-value < 0.01, (***) P-value < 0.001.











