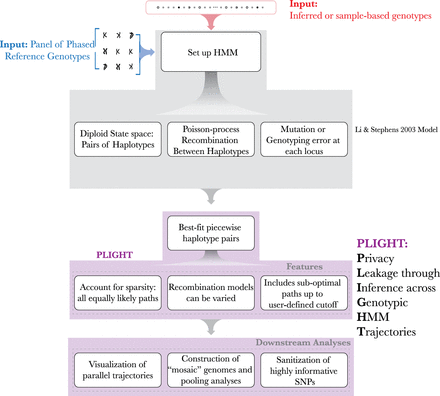
Computational framework of PLIGHT. The inputs of a reference database and of the query SNPs are shown at the top. Next, the population-genetics model by Li and Stephens forms the core of the HMM-based framework. This includes the definition of a diploid state space, a default Poisson process–based recombination model with growth rate proportional to the genomic distance between two loci, and a mutation/error rate at each locus. This model is implemented within PLIGHT, which identifies the best-fit reference haplotype labels in the diploid state space. Special features include the identification of all equally likely trajectories, flexibility in the recombination models used, and an allowance for sub-optimal trajectories to be identified. Furthermore, PLIGHT includes visualization and SNP sanitization modules. The framework also allows for the construction of mosaic genomes and analyses involving the pooling of information across all identified mosaic trajectories.











